Principles

This image represents the initial state of a signaling pathway. The
model is proposed by A. Bugrim
(
A Logic-based Approach for Computational Analysis of Spatially Distributed
Biochemical Networks, Andrej Bugrim, ISMB 2000; see also
this article) and takes into account the
spatial localisation of the molecules. The reaction, diffusion
and transport processes described in the Bugrim model are modeled as
multiset transformations taking place in a nest of multisets. This is
reminiscent of the P system approach.
This figure is automatically generated by the MGS simulation
program. Each box corresponds to a multiset: the external one
represents the universe and contains three elements: the agonist
molecule pictured as a thin cone; the calcium (the thick cone);
and the plasma membrane which is represented as a multiset and figured
by a translucid box. The various molecules anchored in the plasma
membrane are drawn as various solid volumes. The ellipsoidal container
represents the cytosol and the solid sphere in the middle of it, the
nucleus. Such a figure can be generated at each simulation step to
visualize the trajectory of the dynamical system. See below for the
first three steps.

The corresponding MGS program
The organization of the cell is implemented using a nest of multisets and there is a MGS
rule for each chemical reaction and diffusion or transport process. A membrane corresponds
also to a multiset: the multiset of molecules anchored in this membrane. The diagram below gives
a schematic view of the reaction and of the spatial organization involved.
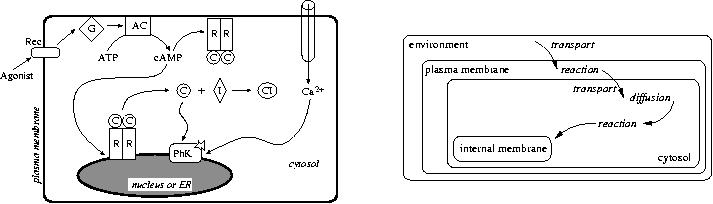
Clément Boin and Nicolas Thibault have developped this program as
part of a student project. Thanks for their work !
- The code of the program (without comment) is available here as a text file.
- The complete program is described (in french) in this technical report (.ps.gz, 992k)
made by Clément Boin and Nicolas Thibault for their TER (a research
project done by students). The first part of this report is devoted to an
application in Artificial Chemistry. The complete code for these two applications
appears in the index.
- The slide presentation of this student project is available as a power-point
file (646k).
- An authorware
presentation of the Bugrim model is available on-line for netscape and explorer browser under windows (we
are sorry but Macromedia does not furnish a plugin for linux).
- And also the same presentation is available as a stand-alone executable (for Windows 9x,
NT, XP). Saves this self-extracting archive. Decompress it just by executing
it and read the README.txt in the directory just created.

this site is
under construction.
Pages started:
May 2002. Last revision: 24 jully 2003.

Pictures, graphics and animations are licensed under a Creative Commons License.
English keywords for indexation:
computer science, programming
language, topological collections, transformation, declarative programming
language, functional languages, simulation of biological processes, cell
model, biological pathway, interaction network, gene regulation, signal
transduction, morphogenesis, developmental biology, integrative simulation,
biological organization, dynamical systems, dynamical structure, Gamma,
CHAM, P system, L system, Paun, Lindenmayer, cellular automata, membrane
computing, aqueous computing, artificial chemistry, GBF, Cayley graph, data
fields, nested collections, rewriting, rule based programming, pattern-matching,
intentional programming, compilation, interpretation, type, type inference,
nested type, polytypism, catamorphism, static analysis, sequence, multiset,
combinatorial algebraic topology, chain complex, chain group.




